The human microbiome has really captured the science spotlight during the last decade, not least through the extensive research undertaken by the National Institutes of Health (NIH). After identifying and cataloging the plethora of micro-organisms that inhabit our mouth, nose, gut, and reproductive tract, the focus of the research has now taken a new direction.
Studies have shown that external factors like diet and environmental exposure, among others, can affect the numbers of the various bacteria that make up our microbiomes. The research is now focused on understanding how microbial population shifts affect three conditions:
- Diabetes
- Inflammatory bowel disease (IBD) and
- Preterm birth
The microbiome and diabetes
We know that insulin resistance is the first step on the road to diabetes. When our systems become less sensitive to the hormone insulin, breaking down sugar and dietary carbohydrates becomes an issue.
70% of individuals with pre-diabetes go on to develop full-blown type-2 diabetes. Insulin resistance, if caught early on, is reversible, but diabetes is less so.
The research shows that the composition of the microbiome of people with insulin resistance is different. These bacteria then cause the immune cells to act in a manner different from people with better insulin sensitivity. As a result, they are more susceptible to disease.
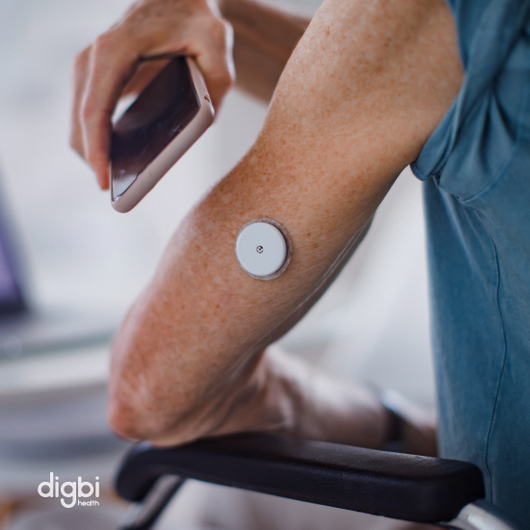
It is possible to predict an "insulin resistance signature" from the microbiome itself. The research, if harnessed to develop a diagnostic test to detect pre-diabetes from stool samples, could mean a drastic reduction in both costs and time compared to current practices.
It is, in fact, already in motion. The regular screening of microbiomes for diabetes could very well become a routine part of a health checkup in the coming years.
The microbiome and IBD
The most prevalent forms of IBD are Crohn’s disease (CD) and ulcerative colitis (UC). They are characterized by debilitating, chronic relapsing and remitting inflammation of the gastrointestinal tract (for CD) or the colon (in UC). These conditions result from a complex interplay among host, microbial, and environmental factors.
The term "flare-up" commonly refers to the onset of the inflammation and accompanying symptoms.
Researchers could identify changes in the types of bacteria that become active during flare-ups compared to during recovery periods. They could also follow the impact on the immune system.
However, the current level of knowledge is not enough to develop, say, a diagnostic test that could predict flare-ups based on a person’s microbiome.
This early work could lay the foundation for further research in this area.
The microbiome and premature birth

Image by Digbi Health
In this case, studying vaginal microbiome has yielded certain microbial profiles linked with a higher incidence of pre-term deliveries. It has been found that women who delivered at less than 37 weeks were less likely to have Lactobacillus crispatus among their vaginal flora, and more likely to contain a diverse community of vaginal microbes, often including four specific types.
The prediction model developed by scientists, incorporating this information improved prediction of premature birth by up to 7% over current methods, and generated more hypotheses to explore.
Understanding the vaginal microbiome might also help to predict IVF success. It can also help establish the most optimal microbiome profile for a healthy pregnancy and full-term delivery.
With the ever-expanding scope of correct diagnoses using microbiome data, testing methods are expected to get even more sophisticated and provide more accurate results.